|
1/26/2024 0 Comments Shine dalgarno sequence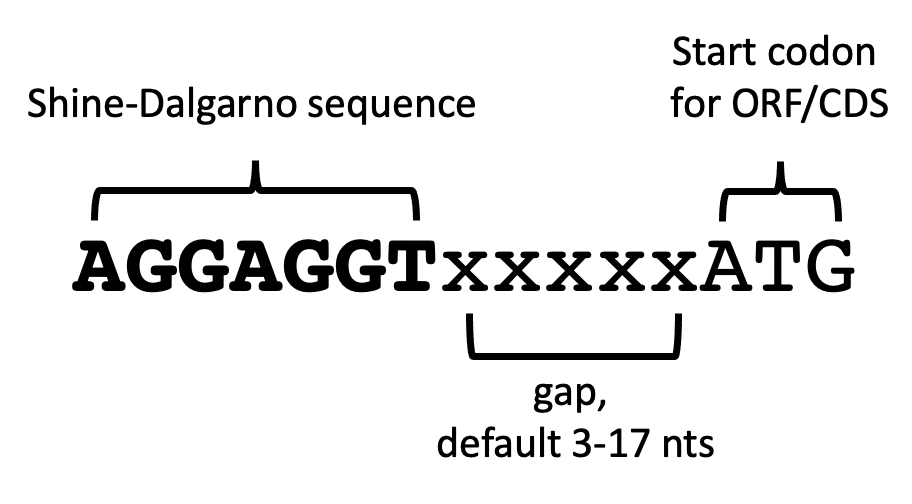
I (inosine) can be found in the anti-codon.įor example, the codons UUU and UUC are both recognized by the tRNA which has GAA in the anti-codon position (making either G - C, or G - U base pairings).Where U, G, A and C can be in either the codon (mRNA) or anti-codon (tRNA).Possible "wobble" codon base pairing (in addition to Watson-Crick): This greater recognition of tRNA is possible due to " wobble" basepair interactions at the third base in the codon/first base in the anti-codon:.We know this is not the case, therefore a single tRNA anti-codon must be able to recognized several different mRNA codon triplets.The Kozak consensus sequence was first identified by Marilyn Kozak in 1984 while she was in the Department of Biological Sciences at the University of Pittsburgh. If perfect Watson-Crick base pairing were required at the codon/anti-codon triplet then 61 different tRNA's would be required. The Shine-Dalgarno sequence, of the prokaryotic RBS, was discovered by John Shine and Lynne Dalgarno in 1975.Class II amino-acyl tRNA synthetases attach their associated amino acids to the tRNA 3' hydroxyl(NOTE: typically hydrophilic amino acids) In this study, we analyzed the correlation between codon usage bias and ShineDalgarno (SD) sequence conservation, using complete genome sequences of.Class I amino-acyl tRNA synthetases attach their associated amino acids to the tRNA 2' hydroxyl (NOTE: typically the hydrophobic amino acids) The Shine-Dalgarno sequence of riboswitch-regulated single mRNAs shows ligand-dependent accessibility bursts.The amino acid is attached at the 3' end of the tRNA to either the 2' hydroxyl or the 3' hydroxyl. Shine Dalgarno Sequence The Shine Dalgarno (SD) grouping is a ribosomal restricting site in bacterial and archaeal courier RNA, by and large, situated around 8.Guanidylate may be methylated at different positions.The SD sequence is located near the start codon which is in contrast to. A Shine-Dalgarno sequence marks the start of each coding sequence, letting the ribosome find the right start codon for each gene. The biggest difference is the existence of the Shine-Dalgarno (SD) sequence in mRNA for bacteria. Determinant of cistron specificity in bacterial. ribose attached to Carbon 5 instead of Nitrogen 1). The scanning mechanism of initiation, which utilizes the Kozak sequence, is found only in eukaryotes and has significant differences from the way bacteria initiate translation. The sequence is complementary to gaucaCCUCCUuaOH at the 3 end of 16S rRNA. A shine Dalgarno sequence is not absolutely essential for translation to occur. Uridylate may be rearranged to produce pseudouridylate (i.e. Shine Dalgarno sequence is immediately upstream of a possible start codon.Uridylate may be methylated to produce Thymidylate.However, after synthesis several bases may be modified: The Shine-Dalgarno (SD) Sequence is a ribosomal binding site in bacterial and archaeal messenger RNA, generally located around 8 bases upstream of the start. tRNA's are synthesized with the standard bases AGCU.Form a series of stem/loop secondary structures.It should be noted, that S1 is only present in Gram-negative bacteria, being absent from Gram-positive species.\) Note that the ribosome does not bind at the AUG start codon, but 5-10 nucleotides upstream. 
What principally attracts ribosome to mRNA initiation region is apparently ribosomal protein S1, which binds to AU-rich sequences found in many prokaryotic mRNAs 15-30 nucleotides upstream of start-codon. The Shine-Dalgarno sequence is thus a ribosome binding site which is necessary for the intiation of translation. Moreover, numerous prokaryotic mRNAs don't possess Shine-Dalgarno sequences at all. In Gram-negative bacteria, however, Shine-Dalgarno sequence presence is not obligatory for ribosome to locate initiator codon, since deletion of Anti-Shine-Dalgarno sequence from 16S rRNA doesn't lead to translation initiation at non-authentic sites. When the Shine-Dalgarno sequence and the anti-Shine-Dalgarno sequence pair, the translation initiation factors IF2-GTP, IF1, IF3, as well as the initiator tRNA fMet-tRNA(fMET) are recruited to the ribosome. This reduction is due to a reduced mRNA-ribosome pairing efficiency, as evidenced by the fact that complementary mutations in the anti-Shine-Dalgarno sequence can restore translation. Mutations in the Shine-Dalgarno sequence can reduce translation. It is a ribosomal binding site in bacterial and archeal mRNA. How to ensure accurate weighing results every day? The Shine-Dalgarno sequence is named after the Australian scientists John Shine and Lynn Dalgarno.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
